Quick Start
Quick start
PhaBOX allows users to copy-and-paste or upload their interesting DNA contigs in FASTA format on the Pipelines pages. When predicting query contigs, PhaBOX server provides two options for analyzing users' sequences:
- Running PhaBOX once for all phage tasks (phage identification, taxa classification, lifestyle prediction, and host prediction).
- Running the subprogram that users are interesting in.
However, when running PhaGCN (taxa classification), PhaTYP (lifestyle prediction), and CHERRY (host prediction) in the single-program mode, users should guarantee their input sequences are all phages. Otherwise, the results may be influenced by the non-phage sequence. In this case, the user can run PhaMer to filter the non-phage genomes first.
Example of input
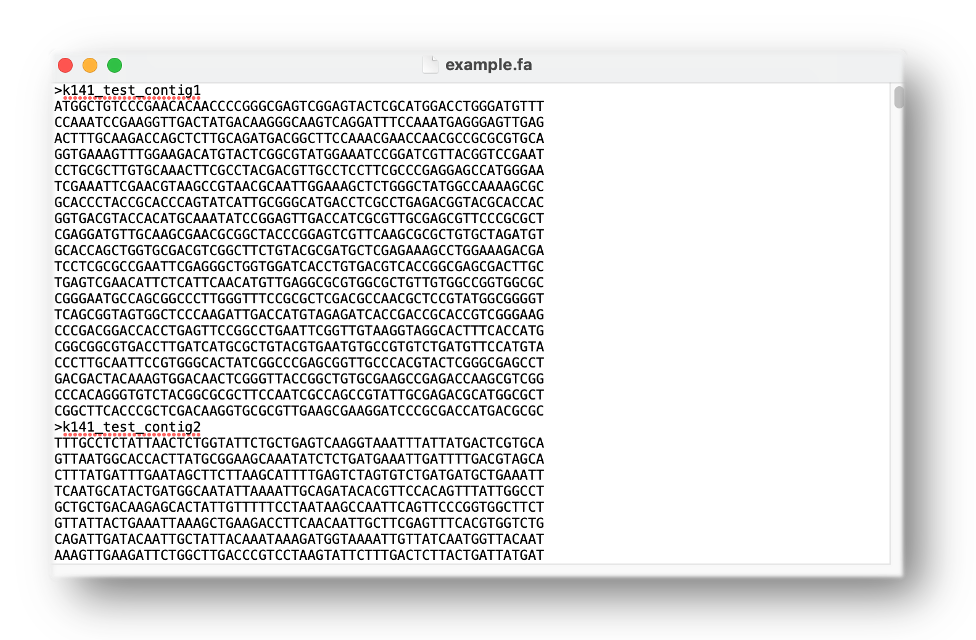
The input of PhaBOX is the standard FASTA file. Each sequence in the FASTA contains a header and DNA sequences. The accession/ID/name of the contigs will be recorded and used in PhaBOX to match the prediction. For example, in the example.fa shown above, the name of the contigs are k141_test_contig1 and k141_test_contig2. The contigs' accession should begin with a letter. Please also make sure there are no special characters in the contigs' accession/ID/name, such as '|', '~', '&', and '$' to avoid unnecessary errors.
Submitting a job
-
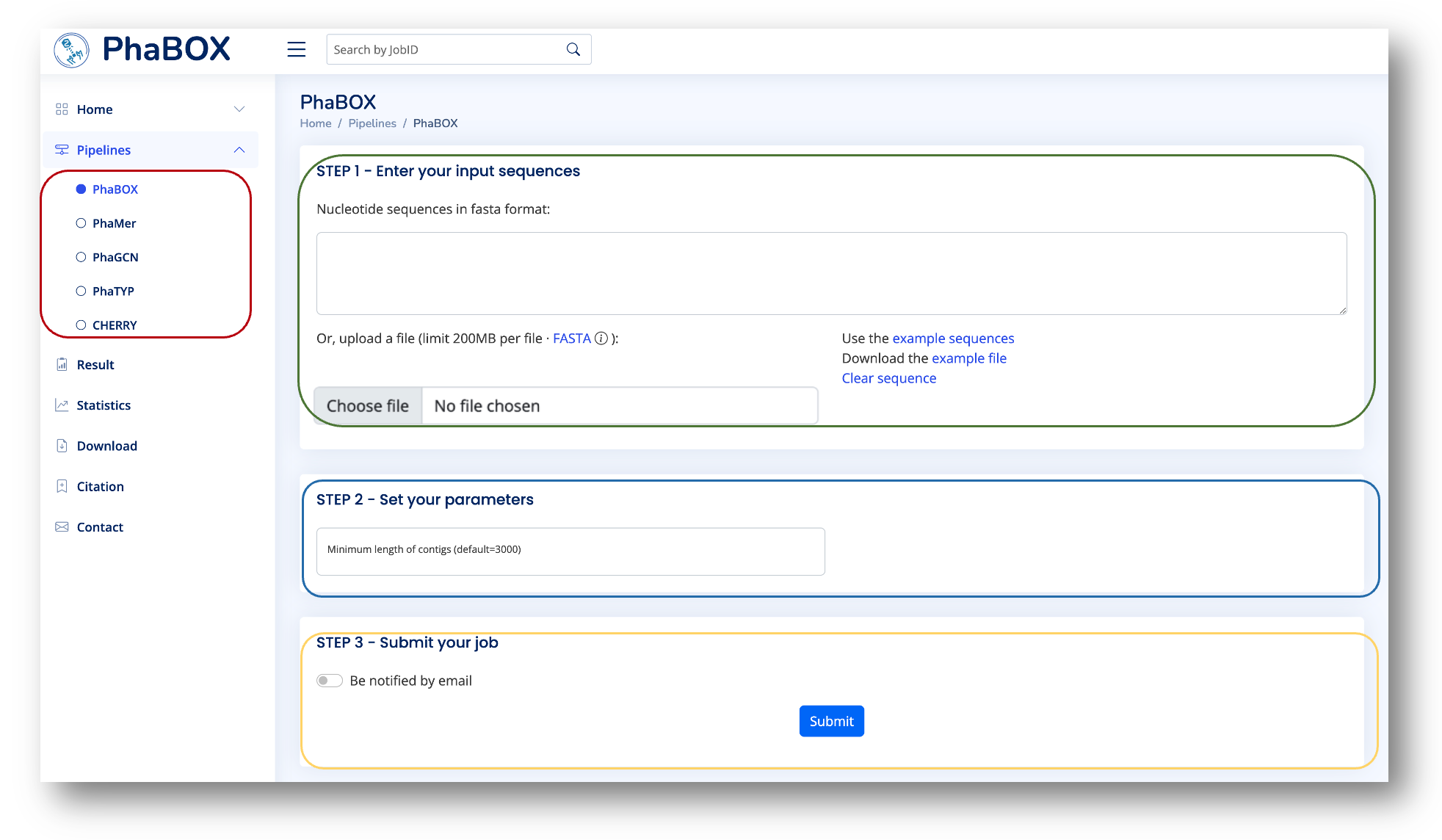
Choose a program you want to run in the red box:
-
PhaMer: phage identification from metagenomic contigs
-
PhaGCN: family-level taxonomy classification
-
PhaTYP: phages' lifestyle prediction
-
CHERRY: Host prediction
-
PhaBOX: run the above tools in one pipeline. It will spend less time compared to run tools seperately.
-
-
Paste or upload your DNA sequences in the green box. The sequences should be in FASTA format. Please click on the example sequence button for instruction.
-
Set a threshold for the minimum length of contigs in the blue box. If you want to use the default parameters, then let it be blank. The program will only handle the contigs longer than the threshold.
-
(Optional) Choose whether you want to be notified by email in the yellow box. If yes, turn on the button and paste your email address. Otherwise, submit your task directly and remember the job ID.
After submission
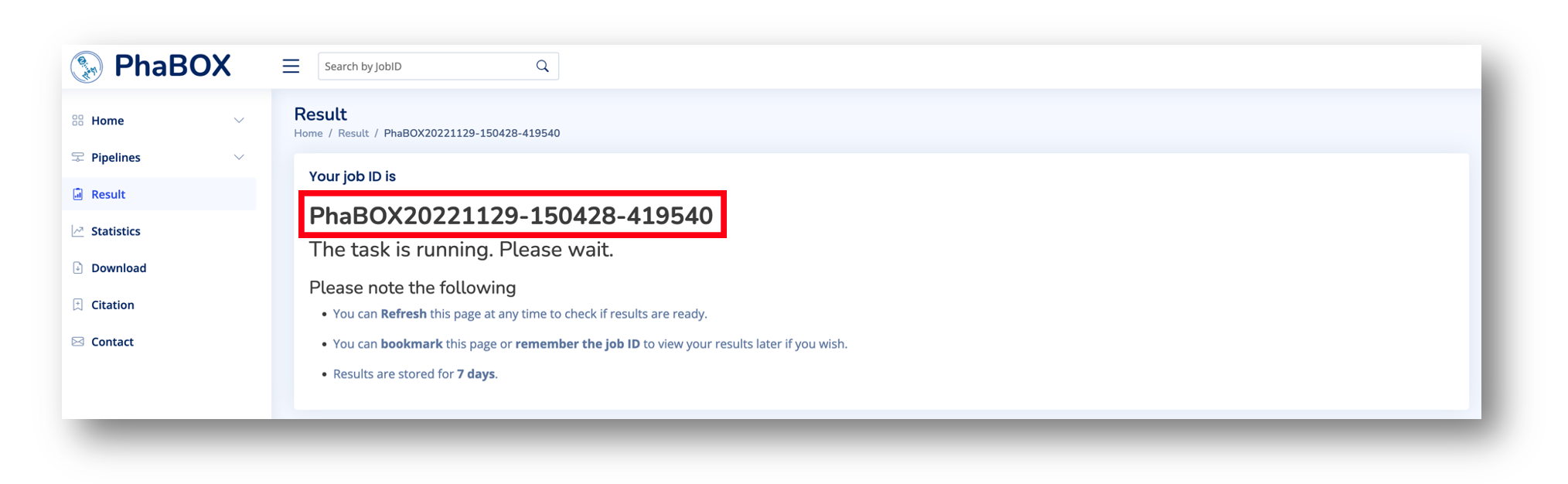
Once you submit your job, the webpage will jump to the Result page and show the job ID of your submission. One example job ID is shown below. Please remember your job ID to find your results.
If you turn on the email notification, you will receive an email once the job is submitted successfully and when the job is finished. An example is shown as below:
- After submitting the job:

- When the job is finished:

The email address used for notification in PhaBOX is: phage.cityu@gmail.com. If you did not receive our email after submitting the job, you may need to check whether it is in your junk mail box. Please add the email into the whitelist if you wish to receive our notification.
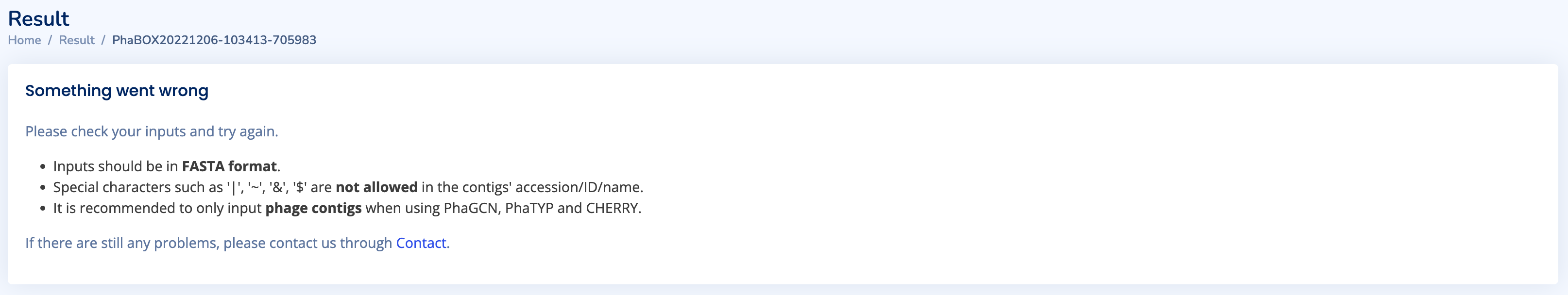
An exception might occur if your upload files are not in FASTA format. The page will return “Please try again" message (as shown below). Please check whether the format of your file is correct. If you have no idea about your file. You can contact us using the Contact function with your jobID.